Rogue Proteins May Underlie Some ALS and Frontotemporal Dementia Cases, Says Study
Discovery May Lead to New Ways of Diagnosing and Treating Both Diseases
ALS—amyotrophic lateral sclerosis—is a neurodegenerative disease that attacks motor neurons in the brain and spinal cord, slowly robbing its victims of their ability to walk, talk, breathe and swallow. In a cruel twist, some ALS patients also develop frontotemporal dementia, a disease that destroys an entirely different set of brain cells—cortical neurons—leading to personality changes, among other effects.
Inherited forms of both diseases have been traced to gene mutations associated with an abnormal buildup of the RNA-binding protein, TDP-43, in the brain. Now, in a study in the journal eLife, Columbia University and New York Genome Center researchers show that TDP-43 and at least three other RNA-binding proteins appear to run similarly amok in ALS and dementia patients without the mutation. Their finding suggests that the dominant form of hereditary ALS and frontotemporal dementia may share molecular underpinnings with far more common versions of ALS and dementia that have no known genetic basis.
“It turns out that if you analyze the biochemical properties of RNA-binding proteins you see it’s not just TDP-43 but several others that are also perturbed,” says Aaron Gitler, a genetics professor at Stanford University who discusses the study in the same issue of eLife. “This is a new concept in how we think about these diseases–not just as TDP-43 diseases, but as RNA-binding protein diseases.”
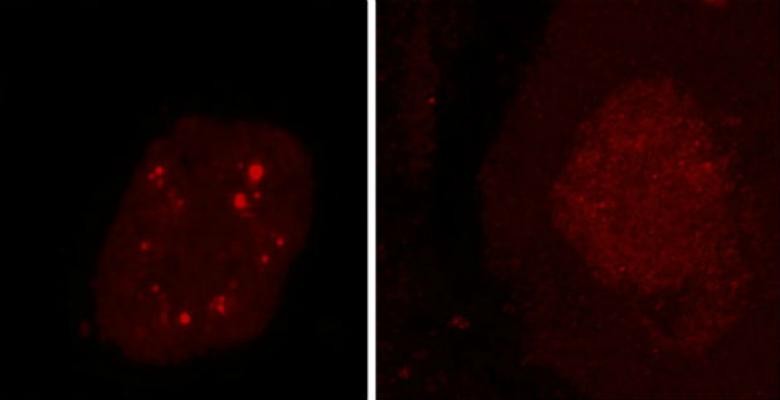
Normally, the TDP-43 protein helps control the expression of messenger RNA, and thus, gene behavior, from inside the cell nucleus. But in the brain cells of nearly all ALS and half of frontotemporal dementia patients, the protein accumulates outside the nucleus, eventually forming clumps big enough to see under a microscope. What causes the buildup, however, remains unclear.
In a pair of landmark studies in Neuron in 2011, researchers identified the genetic mutation responsible for most cases of inherited ALS and frontotemporal dementia. On a small section of chromosome 9, the mutation appears to cause a DNA fragment that normally recurs a handful of times to repeat itself hundreds to thousands of times. Researchers hypothesized that RNA-binding proteins like TDP-43, designed to stick to themselves and similar proteins, would glom on to the repeats and abandon their gene-expression role, eventually causing brain cells to degenerate and die.
Read the Full Study
In a 2016 study in eLife, a team led by Erin Conlon, then a Columbia graduate student, showed that the mutation triggered another RNA-binding protein, called hnRNP H, to form similar clumps that disrupted gene expression. Unlike TDP-43, these clumps were largely invisible to the eye but could be measured with a biochemical test.
On a hunch that a similar pattern might show up in mutation-free patients, who represent a majority of ALS and frontotemporal dementia cases, Conlon and her colleagues in the current study analyzed the brains of 50 people who had died with one or both diseases. To their surprise, in more than half of the brains, they found large amounts of biochemically insoluble hnRNP H and three other RNA-binding proteins—TDP-43, FUS and hnRNP A1—indicating all had stopped regulating gene expression.
“RNA-binding proteins control how much proteins a gene makes,” says the study’s senior author, James Manley, a molecular biology professor at Columbia. “This process goes seriously awry when these aggregates are around.”
Many questions remain, including what causes RNA-binding proteins, in the absence of a mutation, to go haywire in the first place. Another mystery is why the impaired proteins surface in motor and cortical neurons while other brain cells are spared.
Neurons may be especially vulnerable to toxic protein accumulations, because unlike other cells, they can’t divide or be replaced, but that doesn’t explain why motor and cortical neurons are especially sensitive, says Conlon, now a postdoctoral researcher at Rockefeller University.
Though much more still needs to be worked out, the study suggests that a blood test or other non-invasive way to detect ALS and frontotemporal dementia could soon be within reach. The findings also indicate that ALS comes in at least two forms — one in which RNA-binding function is disrupted, and the other with a still-unknown mechanism.
“Our data suggests that we may be able to distinguish one subpopulation of ALS patients from the other,” says study coauthor Dr. Neil Shneider, a neurology professor who heads Columbia University Irving Medical Center’s Eleanor and Lou Gehrig ALS Center. “This could lead to a therapeutic intervention that works selectively on those patients.”
Other authors of the study are Julia Davis-Porada, Columbia; and Delphine Fagegaltier, Phaedra Agius, James Gregory, Isabel Hubbard, Kristy Kang, Duyang Kim and Hemali Phatnani, New York Genome Center. Funding was provided by the National Institutes of Health, ALS Association, Tow Foundation and Eleanor and Lou Gehrig ALS Center.